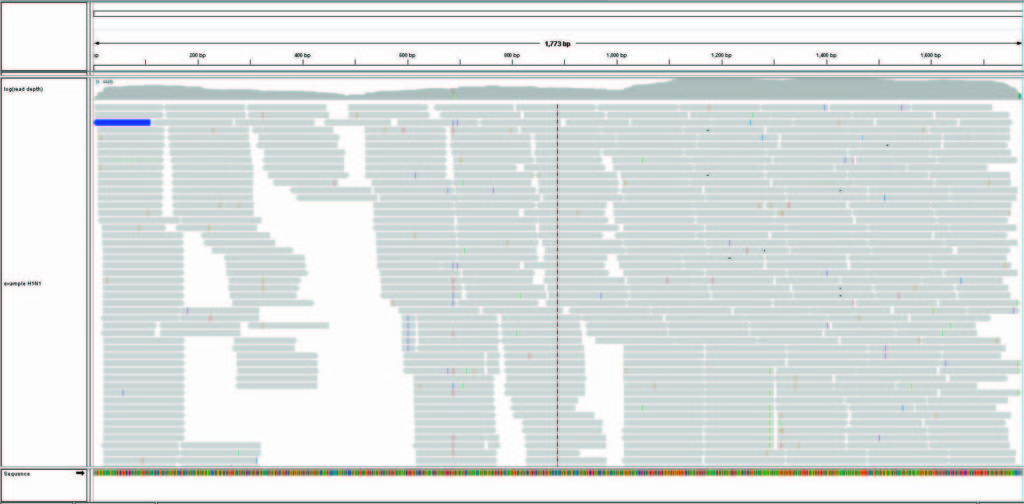
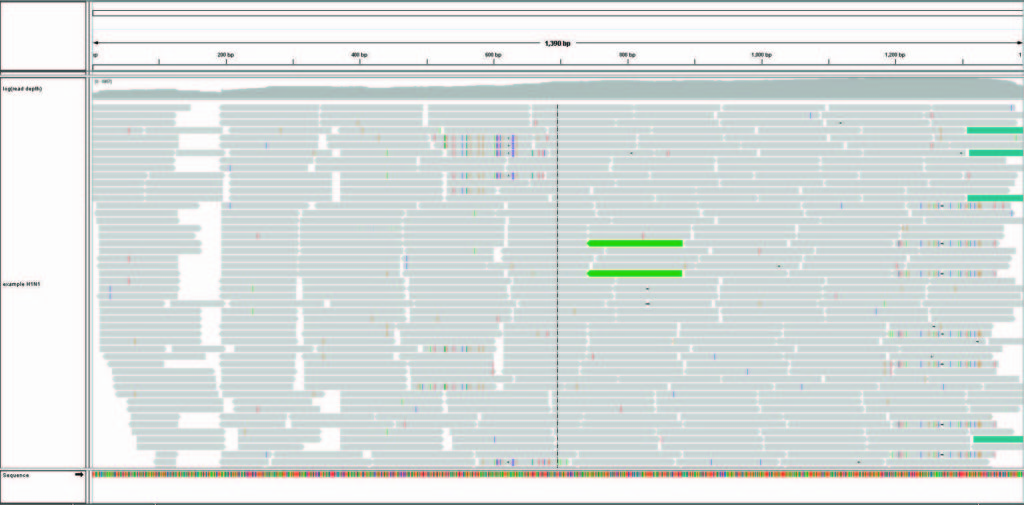
The ONE™ Influenza Sentinel assay was applied on H1N1 influenza virus-positive samples. Example data demonstrates the assay’s capability to (1) identify influenza type, subtype, and strain correctly, (2) recover the entire sequences of HA and NA, and (3) reveal mutations and quasi-species.
References
.
